I lead the Generate group in the Marks lab at Harvard Medical School. We develop protein foundation models, create large-scale benchmarks for robust in silico model evaluation, and apply these models to design novel enzymes with desired properties in collaboration with experimental laboratories.
Protein Foundation Models
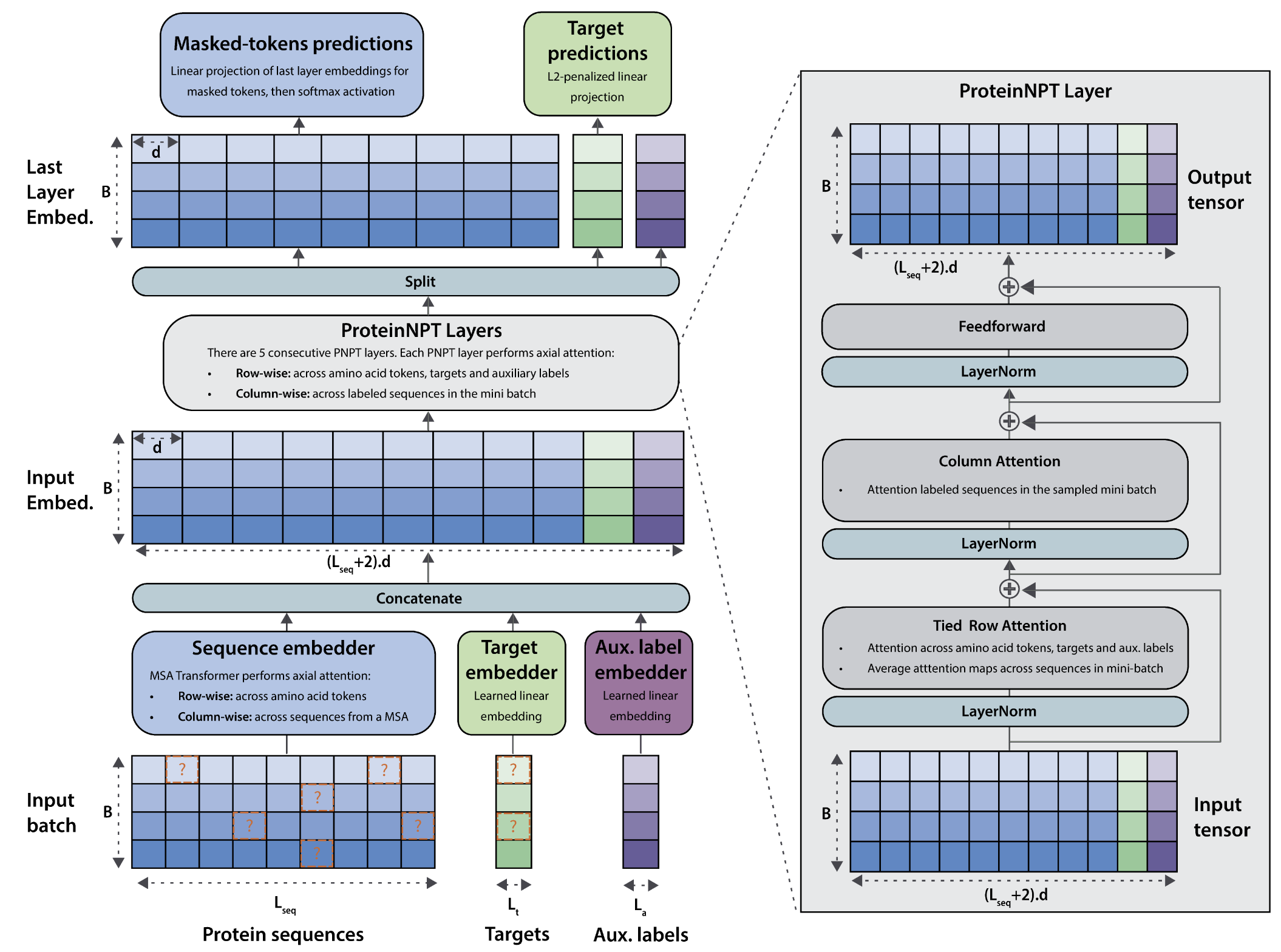
We pioneer generative AI models to learn complex protein fitness landscapes. Our current focus is on multimodal & retrieval-augmented protein models, conditional protein design, advanced sampling, and methods for data-efficient transfer learning across landscapes.
Large-scale Model Evaluation
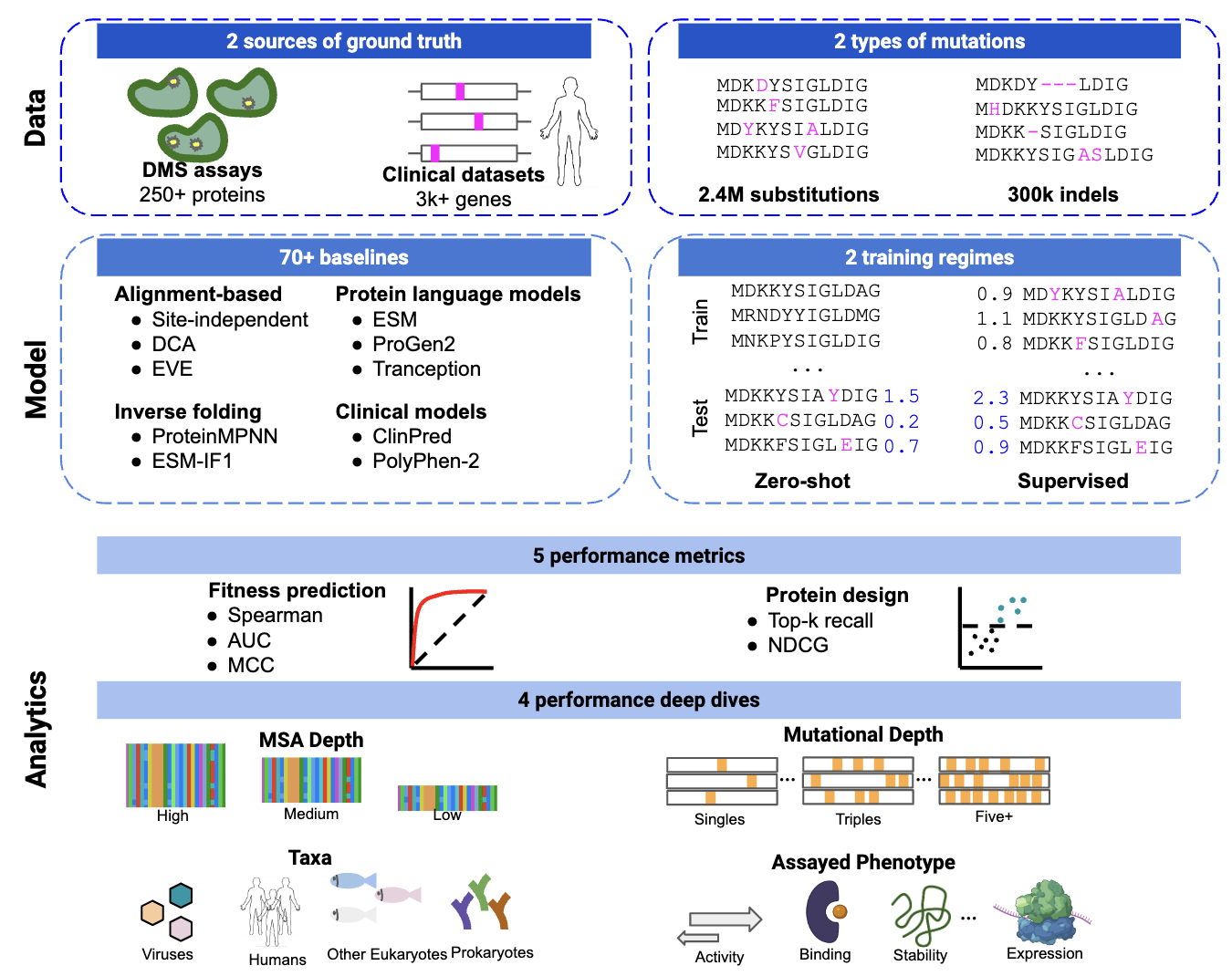
We develop rigorous benchmarking frameworks that set standards for in silico model evaluation, such as ProteinGym and RNAGym. Our benchmarks are based on large-scale high-throughput experiments to promote robust performance in real-world applications.
Applied Enzyme Design
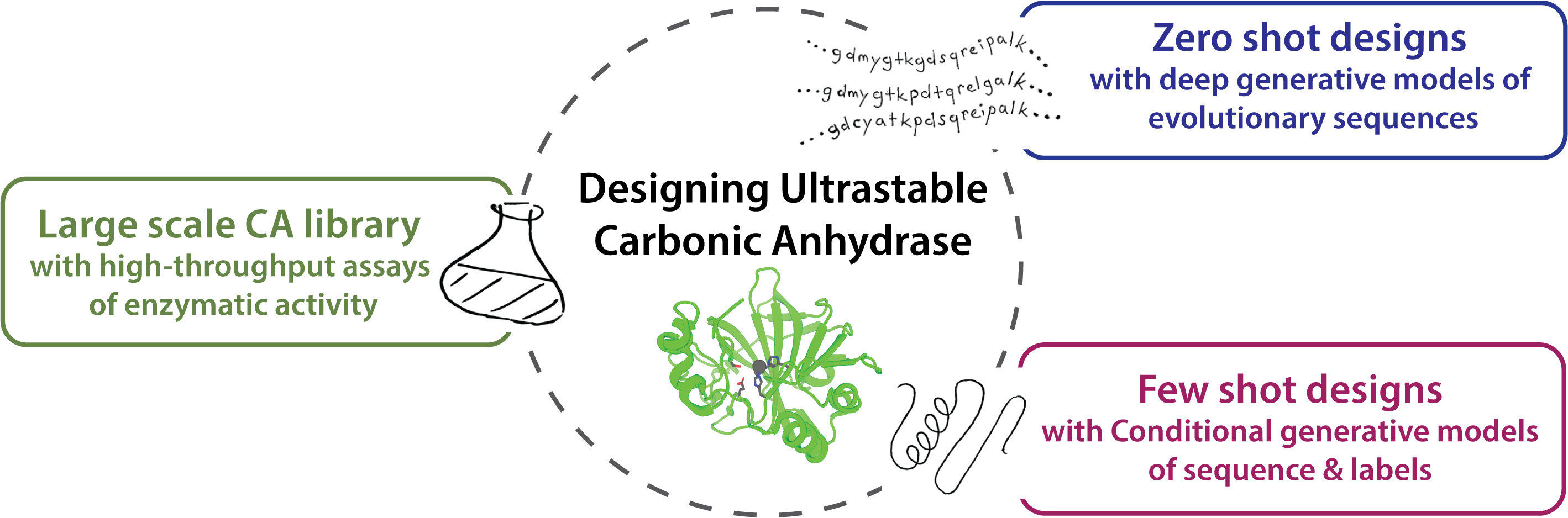
We combine our AI foundation models with high and low-throughput experimental methods to characterize and predict enzyme function, or to design novel enzymes with improved catalytic efficiency or stability for targeted applications in healthcare and sustainability.